Pawel Foszner
POLCOVID: a multicenter multiclass chest X-ray database
Dec 15, 2022Abstract:The outbreak of the SARS-CoV-2 pandemic has put healthcare systems worldwide to their limits, resulting in increased waiting time for diagnosis and required medical assistance. With chest radiographs (CXR) being one of the most common COVID-19 diagnosis methods, many artificial intelligence tools for image-based COVID-19 detection have been developed, often trained on a small number of images from COVID-19-positive patients. Thus, the need for high-quality and well-annotated CXR image databases increased. This paper introduces POLCOVID dataset, containing chest X-ray (CXR) images of patients with COVID-19 or other-type pneumonia, and healthy individuals gathered from 15 Polish hospitals. The original radiographs are accompanied by the preprocessed images limited to the lung area and the corresponding lung masks obtained with the segmentation model. Moreover, the manually created lung masks are provided for a part of POLCOVID dataset and the other four publicly available CXR image collections. POLCOVID dataset can help in pneumonia or COVID-19 diagnosis, while the set of matched images and lung masks may serve for the development of lung segmentation solutions.
CIRCA: comprehensible online system in support of chest X-rays-based COVID-19 diagnosis
Oct 11, 2022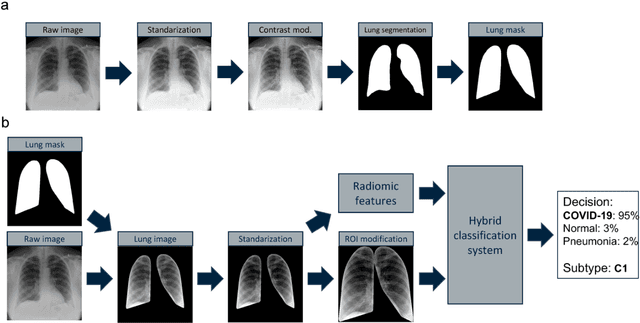
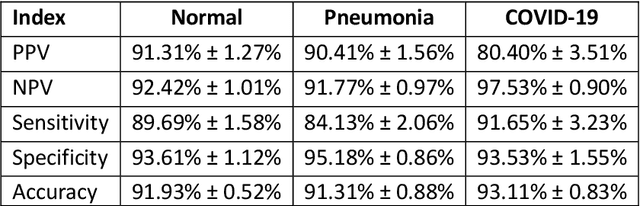
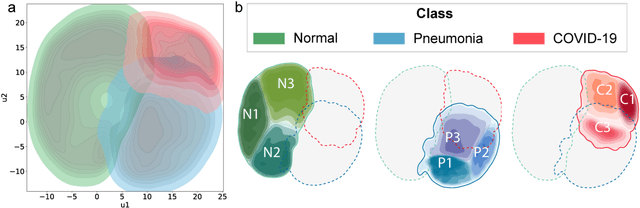
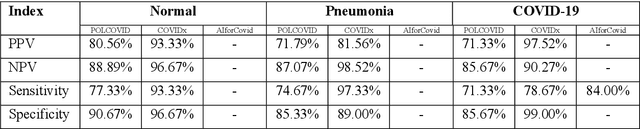
Abstract:Due to the large accumulation of patients requiring hospitalization, the COVID-19 pandemic disease caused a high overload of health systems, even in developed countries. Deep learning techniques based on medical imaging data can help in the faster detection of COVID-19 cases and monitoring of disease progression. Regardless of the numerous proposed solutions for lung X-rays, none of them is a product that can be used in the clinic. Five different datasets (POLCOVID, AIforCOVID, COVIDx, NIH, and artificially generated data) were used to construct a representative dataset of 23 799 CXRs for model training; 1 050 images were used as a hold-out test set, and 44 247 as independent test set (BIMCV database). A U-Net-based model was developed to identify a clinically relevant region of the CXR. Each image class (normal, pneumonia, and COVID-19) was divided into 3 subtypes using a 2D Gaussian mixture model. A decision tree was used to aggregate predictions from the InceptionV3 network based on processed CXRs and a dense neural network on radiomic features. The lung segmentation model gave the Sorensen-Dice coefficient of 94.86% in the validation dataset, and 93.36% in the testing dataset. In 5-fold cross-validation, the accuracy for all classes ranged from 91% to 93%, keeping slightly higher specificity than sensitivity and NPV than PPV. In the hold-out test set, the balanced accuracy ranged between 68% and 100%. The highest performance was obtained for the subtypes N1, P1, and C1. A similar performance was obtained on the independent dataset for normal and COVID-19 class subtypes. Seventy-six percent of COVID-19 patients wrongly classified as normal cases were annotated by radiologists as with no signs of disease. Finally, we developed the online service (https://circa.aei.polsl.pl) to provide access to fast diagnosis support tools.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge