Betül Güvenç Paltun
Robust Intrusion Detection System with Explainable Artificial Intelligence
Mar 07, 2025Abstract:Machine learning (ML) models serve as powerful tools for threat detection and mitigation; however, they also introduce potential new risks. Adversarial input can exploit these models through standard interfaces, thus creating new attack pathways that threaten critical network operations. As ML advancements progress, adversarial strategies become more advanced, and conventional defenses such as adversarial training are costly in computational terms and often fail to provide real-time detection. These methods typically require a balance between robustness and model performance, which presents challenges for applications that demand instant response. To further investigate this vulnerability, we suggest a novel strategy for detecting and mitigating adversarial attacks using eXplainable Artificial Intelligence (XAI). This approach is evaluated in real time within intrusion detection systems (IDS), leading to the development of a zero-touch mitigation strategy. Additionally, we explore various scenarios in the Radio Resource Control (RRC) layer within the Open Radio Access Network (O-RAN) framework, emphasizing the critical need for enhanced mitigation techniques to strengthen IDS defenses against advanced threats and implement a zero-touch mitigation solution. Extensive testing across different scenarios in the RRC layer of the O-RAN infrastructure validates the ability of the framework to detect and counteract integrated RRC-layer attacks when paired with adversarial strategies, emphasizing the essential need for robust defensive mechanisms to strengthen IDS against complex threats.
DIVERSE: bayesian Data IntegratiVE learning for precise drug ResponSE prediction
Mar 31, 2021

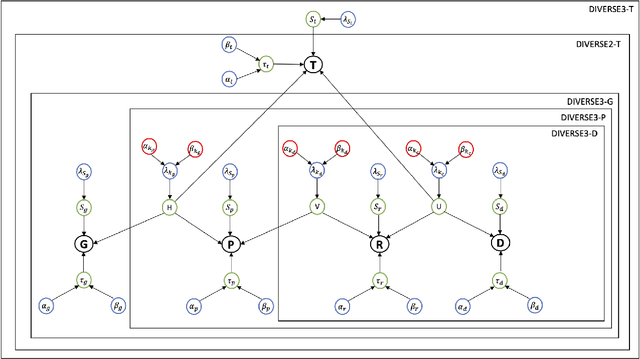
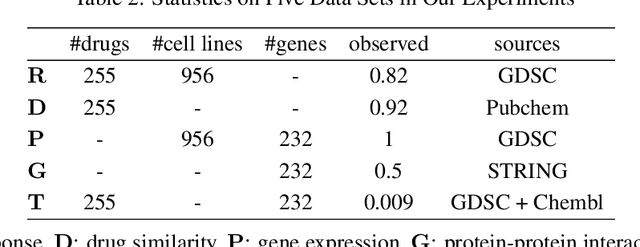
Abstract:Detecting predictive biomarkers from multi-omics data is important for precision medicine, to improve diagnostics of complex diseases and for better treatments. This needs substantial experimental efforts that are made difficult by the heterogeneity of cell lines and huge cost. An effective solution is to build a computational model over the diverse omics data, including genomic, molecular, and environmental information. However, choosing informative and reliable data sources from among the different types of data is a challenging problem. We propose DIVERSE, a framework of Bayesian importance-weighted tri- and bi-matrix factorization(DIVERSE3 or DIVERSE2) to predict drug responses from data of cell lines, drugs, and gene interactions. DIVERSE integrates the data sources systematically, in a step-wise manner, examining the importance of each added data set in turn. More specifically, we sequentially integrate five different data sets, which have not all been combined in earlier bioinformatic methods for predicting drug responses. Empirical experiments show that DIVERSE clearly outperformed five other methods including three state-of-the-art approaches, under cross-validation, particularly in out-of-matrix prediction, which is closer to the setting of real use cases and more challenging than simpler in-matrix prediction. Additionally, case studies for discovering new drugs further confirmed the performance advantage of DIVERSE.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge