Bargava Subramanian
A Multi-Site Study on AI-Driven Pathology Detection and Osteoarthritis Grading from Knee X-Ray
Mar 28, 2025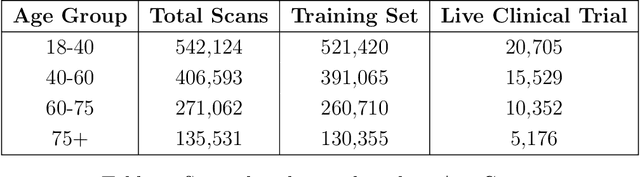


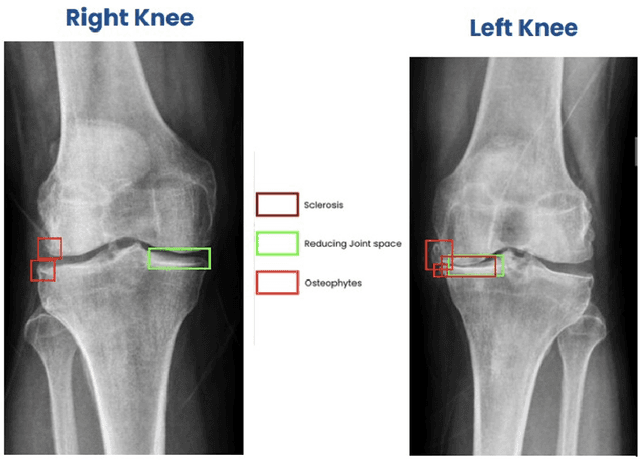
Abstract:Introduction: Bone health disorders like osteoarthritis and osteoporosis pose major global health challenges, often leading to delayed diagnoses due to limited diagnostic tools. This study presents an AI-powered system that analyzes knee X-rays to detect key pathologies, including joint space narrowing, sclerosis, osteophytes, tibial spikes, alignment issues, and soft tissue anomalies. It also grades osteoarthritis severity, enabling timely, personalized treatment. Study Design: The research used 1.3 million knee X-rays from a multi-site Indian clinical trial across government, private, and SME hospitals. The dataset ensured diversity in demographics, imaging equipment, and clinical settings. Rigorous annotation and preprocessing yielded high-quality training datasets for pathology-specific models like ResNet15 for joint space narrowing and DenseNet for osteoarthritis grading. Performance: The AI system achieved strong diagnostic accuracy across diverse imaging environments. Pathology-specific models excelled in precision, recall, and NPV, validated using Mean Squared Error (MSE), Intersection over Union (IoU), and Dice coefficient. Subgroup analyses across age, gender, and manufacturer variations confirmed generalizability for real-world applications. Conclusion: This scalable, cost-effective solution for bone health diagnostics demonstrated robust performance in a multi-site trial. It holds promise for widespread adoption, especially in resource-limited healthcare settings, transforming bone health management and enabling proactive patient care.
AI-Driven MRI Spine Pathology Detection: A Comprehensive Deep Learning Approach for Automated Diagnosis in Diverse Clinical Settings
Mar 26, 2025



Abstract:Study Design This study presents the development of an autonomous AI system for MRI spine pathology detection, trained on a dataset of 2 million MRI spine scans sourced from diverse healthcare facilities across India. The AI system integrates advanced architectures, including Vision Transformers, U-Net with cross-attention, MedSAM, and Cascade R-CNN, enabling comprehensive classification, segmentation, and detection of 43 distinct spinal pathologies. The dataset is balanced across age groups, genders, and scanner manufacturers to ensure robustness and adaptability. Subgroup analyses were conducted to validate the model's performance across different patient demographics, imaging conditions, and equipment types. Performance The AI system achieved up to 97.9 percent multi-pathology detection, demonstrating consistent performance across age, gender, and manufacturer subgroups. The normal vs. abnormal classification achieved 98.0 percent accuracy, and the system was deployed across 13 major healthcare enterprises in India, encompassing diagnostic centers, large hospitals, and government facilities. During deployment, it processed approximately 100,000 plus MRI spine scans, leading to reduced reporting times and increased diagnostic efficiency by automating the identification of common spinal conditions. Conclusion The AI system's high precision and recall validate its capability as a reliable tool for autonomous normal/abnormal classification, pathology segmentation, and detection. Its scalability and adaptability address critical diagnostic gaps, optimize radiology workflows, and improve patient care across varied healthcare environments in India.
3D Convolutional Neural Networks for Improved Detection of Intracranial bleeding in CT Imaging
Mar 26, 2025

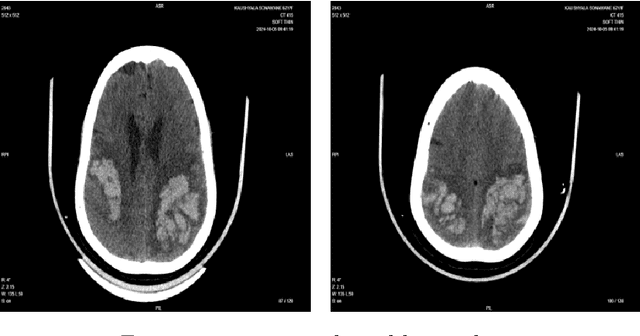

Abstract:Background: Intracranial bleeding (IB) is a life-threatening condition caused by traumatic brain injuries, including epidural, subdural, subarachnoid, and intraparenchymal hemorrhages. Rapid and accurate detection is crucial to prevent severe complications. Traditional imaging can be slow and prone to variability, especially in high-pressure scenarios. Artificial Intelligence (AI) provides a solution by quickly analyzing medical images, identifying subtle hemorrhages, and flagging urgent cases. By enhancing diagnostic speed and accuracy, AI improves workflows and patient care. This article explores AI's role in transforming IB detection in emergency settings. Methods: A U-shaped 3D Convolutional Neural Network (CNN) automates IB detection and classification in volumetric CT scans. Advanced preprocessing, including CLAHE and intensity normalization, enhances image quality. The architecture preserves spatial and contextual details for precise segmentation. A dataset of 2,912 annotated CT scans was used for training and evaluation. Results: The model achieved high performance across major bleed types, with precision, recall, and accuracy exceeding 90 percent in most cases 96 percent precision for epidural hemorrhages and 94 percent accuracy for subarachnoid hemorrhages. Its ability to classify and localize hemorrhages highlights its clinical reliability. Conclusion: This U-shaped 3D CNN offers a scalable solution for automating IB detection, reducing diagnostic delays, and improving emergency care outcomes. Future work will expand dataset diversity, optimize real-time processing, and integrate multimodal data for enhanced clinical applicability.
Vision-Language Models for Acute Tuberculosis Diagnosis: A Multimodal Approach Combining Imaging and Clinical Data
Mar 20, 2025

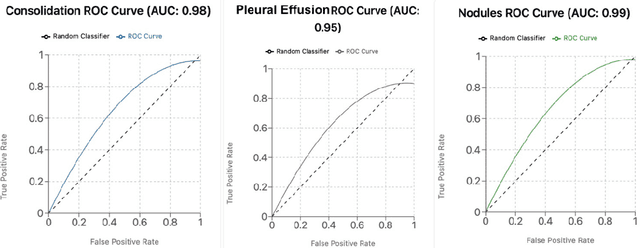
Abstract:Background: This study introduces a Vision-Language Model (VLM) leveraging SIGLIP and Gemma-3b architectures for automated acute tuberculosis (TB) screening. By integrating chest X-ray images and clinical notes, the model aims to enhance diagnostic accuracy and efficiency, particularly in resource-limited settings. Methods: The VLM combines visual data from chest X-rays with clinical context to generate detailed, context-aware diagnostic reports. The architecture employs SIGLIP for visual encoding and Gemma-3b for decoding, ensuring effective representation of acute TB-specific pathologies and clinical insights. Results: Key acute TB pathologies, including consolidation, cavities, and nodules, were detected with high precision (97percent) and recall (96percent). The model demonstrated strong spatial localization capabilities and robustness in distinguishing TB-positive cases, making it a reliable tool for acute TB diagnosis. Conclusion: The multimodal capability of the VLM reduces reliance on radiologists, providing a scalable solution for acute TB screening. Future work will focus on improving the detection of subtle pathologies and addressing dataset biases to enhance its generalizability and application in diverse global healthcare settings.
Advancing Chronic Tuberculosis Diagnostics Using Vision-Language Models: A Multi modal Framework for Precision Analysis
Mar 17, 2025



Abstract:Background This study proposes a Vision-Language Model (VLM) leveraging the SIGLIP encoder and Gemma-3b transformer decoder to enhance automated chronic tuberculosis (TB) screening. By integrating chest X-ray images with clinical data, the model addresses the challenges of manual interpretation, improving diagnostic consistency and accessibility, particularly in resource-constrained settings. Methods The VLM architecture combines a Vision Transformer (ViT) for visual encoding and a transformer-based text encoder to process clinical context, such as patient histories and treatment records. Cross-modal attention mechanisms align radiographic features with textual information, while the Gemma-3b decoder generates comprehensive diagnostic reports. The model was pre-trained on 5 million paired medical images and texts and fine-tuned using 100,000 chronic TB-specific chest X-rays. Results The model demonstrated high precision (94 percent) and recall (94 percent) for detecting key chronic TB pathologies, including fibrosis, calcified granulomas, and bronchiectasis. Area Under the Curve (AUC) scores exceeded 0.93, and Intersection over Union (IoU) values were above 0.91, validating its effectiveness in detecting and localizing TB-related abnormalities. Conclusion The VLM offers a robust and scalable solution for automated chronic TB diagnosis, integrating radiographic and clinical data to deliver actionable and context-aware insights. Future work will address subtle pathologies and dataset biases to enhance the model's generalizability, ensuring equitable performance across diverse populations and healthcare settings.
AI and Deep Learning for Automated Segmentation and Quantitative Measurement of Spinal Structures in MRI
Mar 14, 2025



Abstract:Background: Accurate spinal structure measurement is crucial for assessing spine health and diagnosing conditions like spondylosis, disc herniation, and stenosis. Manual methods for measuring intervertebral disc height and spinal canal diameter are subjective and time-consuming. Automated solutions are needed to improve accuracy, efficiency, and reproducibility in clinical practice. Purpose: This study develops an autonomous AI system for segmenting and measuring key spinal structures in MRI scans, focusing on intervertebral disc height and spinal canal anteroposterior (AP) diameter in the cervical, lumbar, and thoracic regions. The goal is to reduce clinician workload, enhance diagnostic consistency, and improve assessments. Methods: The AI model leverages deep learning architectures, including UNet, nnU-Net, and CNNs. Trained on a large proprietary MRI dataset, it was validated against expert annotations. Performance was evaluated using Dice coefficients and segmentation accuracy. Results: The AI model achieved Dice coefficients of 0.94 for lumbar, 0.91 for cervical, and 0.90 for dorsal spine segmentation (D1-D12). It precisely measured spinal parameters like disc height and canal diameter, demonstrating robustness and clinical applicability. Conclusion: The AI system effectively automates MRI-based spinal measurements, improving accuracy and reducing clinician workload. Its consistent performance across spinal regions supports clinical decision-making, particularly in high-demand settings, enhancing spinal assessments and patient outcomes.
Deep Learning-Based Automated Workflow for Accurate Segmentation and Measurement of Abdominal Organs in CT Scans
Mar 13, 2025Abstract:Background: Automated analysis of CT scans for abdominal organ measurement is crucial for improving diagnostic efficiency and reducing inter-observer variability. Manual segmentation and measurement of organs such as the kidneys, liver, spleen, and prostate are time-consuming and subject to inconsistency, underscoring the need for automated approaches. Purpose: The purpose of this study is to develop and validate an automated workflow for the segmentation and measurement of abdominal organs in CT scans using advanced deep learning models, in order to improve accuracy, reliability, and efficiency in clinical evaluations. Methods: The proposed workflow combines nnU-Net, U-Net++ for organ segmentation, followed by a 3D RCNN model for measuring organ volumes and dimensions. The models were trained and evaluated on CT datasets with metrics such as precision, recall, and Mean Squared Error (MSE) to assess performance. Segmentation quality was verified for its adaptability to variations in patient anatomy and scanner settings. Results: The developed workflow achieved high precision and recall values, exceeding 95 for all targeted organs. The Mean Squared Error (MSE) values were low, indicating a high level of consistency between predicted and ground truth measurements. The segmentation and measurement pipeline demonstrated robust performance, providing accurate delineation and quantification of the kidneys, liver, spleen, and prostate. Conclusion: The proposed approach offers an automated, efficient, and reliable solution for abdominal organ measurement in CT scans. By significantly reducing manual intervention, this workflow enhances measurement accuracy and consistency, with potential for widespread clinical implementation. Future work will focus on expanding the approach to other organs and addressing complex pathological cases.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge