A. S. Boev
Training a discrete variational autoencoder for generative chemistry and drug design on a quantum annealer
Aug 26, 2021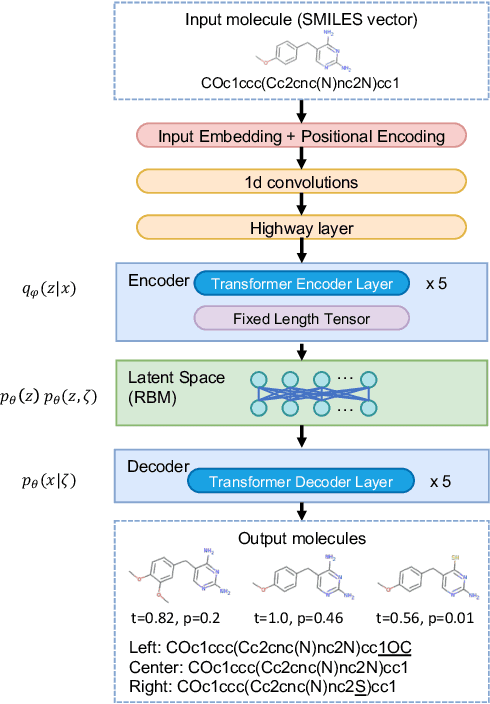



Abstract:Deep generative chemistry models emerge as powerful tools to expedite drug discovery. However, the immense size and complexity of the structural space of all possible drug-like molecules pose significant obstacles, which could be overcome with hybrid architectures combining quantum computers with deep classical networks. We built a compact discrete variational autoencoder (DVAE) with a Restricted Boltzmann Machine (RBM) of reduced size in its latent layer. The size of the proposed model was small enough to fit on a state-of-the-art D-Wave quantum annealer and allowed training on a subset of the ChEMBL dataset of biologically active compounds. Finally, we generated $4290$ novel chemical structures with medicinal chemistry and synthetic accessibility properties in the ranges typical for molecules from ChEMBL. The experimental results point towards the feasibility of using already existing quantum annealing devices for drug discovery problems, which opens the way to building quantum generative models for practically relevant applications.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge