DenseUNets with feedback non-local attention for the segmentation of specular microscopy images of the corneal endothelium with guttae
Paper and Code
Mar 21, 2022
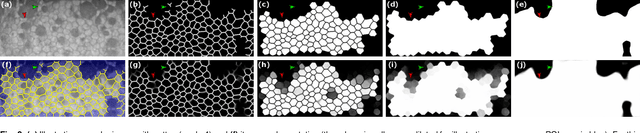
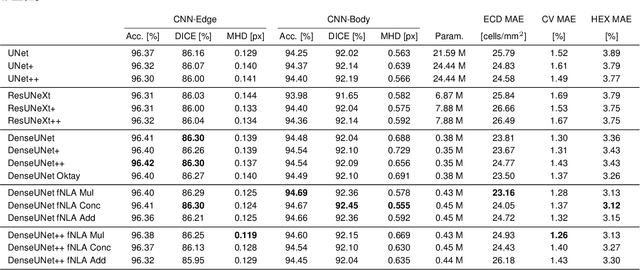

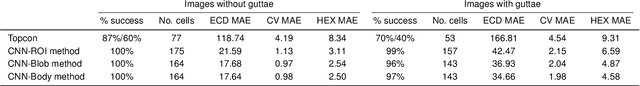
To estimate the corneal endothelial parameters from specular microscopy images depicting cornea guttata (Fuchs dystrophy), we propose a new deep learning methodology that includes a novel attention mechanism named feedback non-local attention (fNLA). Our approach first infers the cell edges, then selects the cells that are well detected, and finally applies a postprocessing method to correct mistakes and provide the binary segmentation from which the corneal parameters are estimated (cell density [ECD], coefficient of variation [CV], and hexagonality [HEX]). In this study, we analyzed 1203 images acquired with a Topcon SP-1P microscope, 500 of which contained guttae. Manual segmentation was performed in all images. We compared the results of different networks (UNet, ResUNeXt, DenseUNets, UNet++) and found that DenseUNets with fNLA provided the best performance, with a mean absolute error of 23.16 [cells/mm$^{2}$] in ECD, 1.28 [%] in CV, and 3.13 [%] in HEX, which was 3-6 times smaller than the error obtained by Topcon's built-in software. Our approach handled the cells affected by guttae remarkably well, detecting cell edges occluded by small guttae while discarding areas covered by large guttae. Overall, the proposed method obtained accurate estimations in extremely challenging specular images.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge